Teresa Carvalho and Ricardo Quiteres wrap up Masters thesis work
November 08, 2019
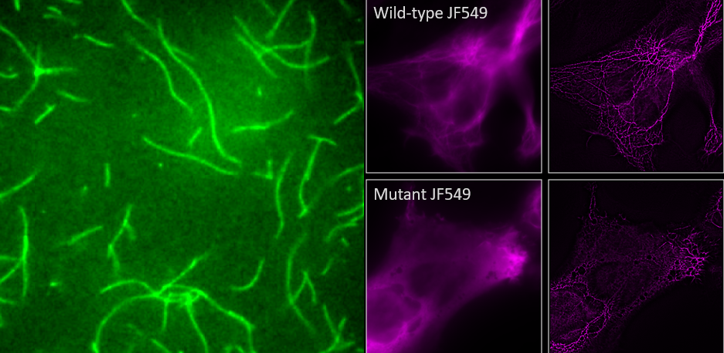
Congrats and good luck to Teresa Carvalho and Ricardo Quiteres who submitted their Masters theses in the last two months and will be defending in the next month! Teresa studied how bacterial effectors influence actin filamentation with a combination of in vitro and in vivo approaches (pictured left) and Ricardo studied intermediate filament defects associated with a neurodegenerative disorder by superresolution microscopy (pictured right).
view this post on its own page
New paper published in mSphere
May 29, 2019
João Silva’s Masters research culimated in a paper published in mSphere today: Plasmids for Independently Tunable, Low-Noise Expression of Two Genes. The paper reports the development and characterization of two compatible plasmids that we co-transformed into E. coli to independently induce GFP (GFPmut2) and RFP (mScarlet-I) expression. Together with Soraia Lopes and Diogo Grilo in the lab, we also showed how the new IPTG-inducible plasmid that João created can be used for imaging single mRNA molecules.
view this post on its own page
Goodbye to Soraia Lopes
March 31, 2019
After almost a year and a half, Soraia’s time in our lab came to an end this week. She was instrumental in getting the lab off the ground and contributed to many on-going projects in RNA detection. She’s off to work in regenerative medicine approaches to ALS treatment!
view this post on its own page
Open source microscope laser/led illumination plans posted
March 21, 2019
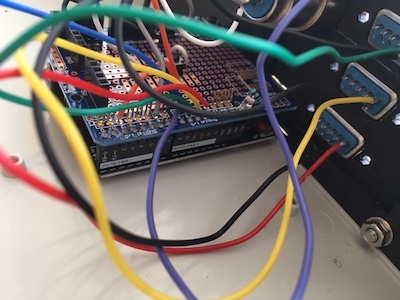

We’ve upgraded our microscopy setup to add LED illumination and allow for rapid modulation of 5 laser lines using Micro-Manager. Plans to reproduce what we’ve built are available at the github repo.

Our design builds on several other free/open components that are described in github readme.
view this post on its own page
New manuscript posted to bioRxiv
January 09, 2019
Congratulations to João Silva and everyone else in the lab who contributed to the first manuscript arising from his masters work. The manuscript, “Plasmids for independently tunable, low-noise expression of two genes,” is now available on bioRxiv. We characterized new plasmids that can be combined to tune the expression of two different genes in E. coli, and we discuss how these plasmids can be used to improve a variety of microbiology experiments. We also showed some data for single-molecule mRNA detection using the bacteriophage PP7 coat protein. The figure below shows how we can use this system to change the expression levels of GFP and mScarlet-I independently.
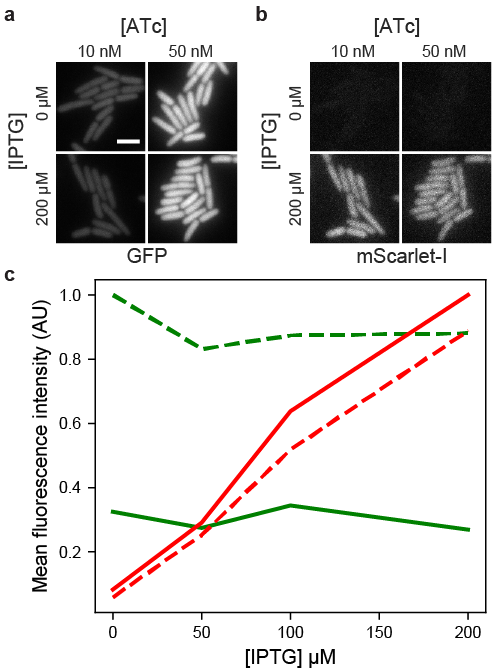
The manuscript was written using Manubot, a revolutionary tool for composing scientific manuscripts online.
view this post on its own page




